Copyright 2021-2023 Lawrence Livermore National Security, LLC and other MuyGPyS Project Developers. See the top-level COPYRIGHT file for details.
SPDX-License-Identifier: MIT
Fast Posterior Mean Tutorial
This notebook walks through the fast posterior mean workflow presented in Fast Gaussian Process Posterior Mean Prediction via Local Cross Validation and Precomputation (Dunton et. al 2022) and explains the relevant components of MuyGPyS.
The cell below uses the same code as that found in univariate_regression_tutorial.ipynb. This includes generating the synthetic data from a GP and training two MuyGPs models to fit the data using Bayesian optimization.
[2]:
import matplotlib.pyplot as plt
import numpy as np
from utils import UnivariateSampler
from MuyGPyS._test.gp import benchmark_sample, BenchmarkGP
from MuyGPyS.gp.distortion import IsotropicDistortion, NullDistortion, l2
from MuyGPyS.gp.hyperparameter import ScalarHyperparameter
from MuyGPyS.gp.kernels import Matern
from MuyGPyS.gp.noise import HomoscedasticNoise
np.random.seed(0)
lb = -10.0
ub = 10.0
data_count = 5001
train_step = 10
nugget_var = 1e-14
fixed_length_scale = 1.0
measurement_eps = 1e-3
sampler = UnivariateSampler(
lb=lb,
ub=ub,
data_count=data_count,
train_step=train_step,
kernel=Matern(
nu=ScalarHyperparameter(0.5),
metric=IsotropicDistortion(
l2,
length_scale=ScalarHyperparameter(fixed_length_scale),
),
),
eps=HomoscedasticNoise(nugget_var),
measurement_eps=HomoscedasticNoise(measurement_eps),
)
train_features, test_features = sampler.features()
test_count, _ = test_features.shape
train_count, _ = train_features.shape
train_responses, test_responses = sampler.sample()
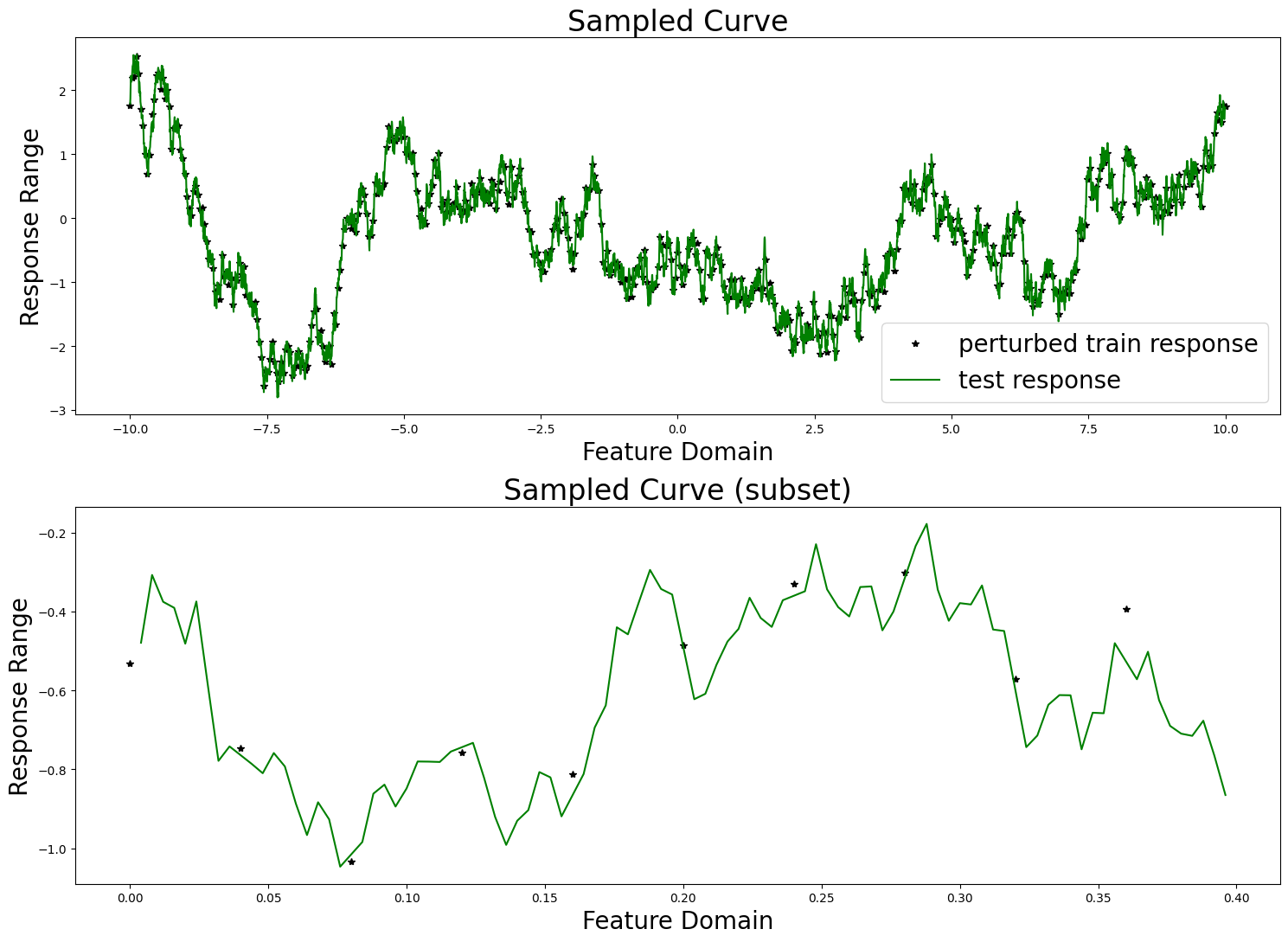
sampler.plot_sample()
from MuyGPyS.neighbors import NN_Wrapper
nn_count = 10
nbrs_lookup = NN_Wrapper(train_features, nn_count, nn_method="exact",algorithm="ball_tree")
from MuyGPyS.optimize.batch import sample_batch
batch_count = train_count
batch_indices, batch_nn_indices = sample_batch(
nbrs_lookup, batch_count, train_count
)
from MuyGPyS.gp import MuyGPS
muygps = MuyGPS(
kernel=Matern(
nu=ScalarHyperparameter(0.5),
metric=IsotropicDistortion(
metric=l2,
length_scale=ScalarHyperparameter(
"log_sample", (0.1, 5.0))
),
),
eps=HomoscedasticNoise(measurement_eps),
)
from MuyGPyS.gp.tensors import crosswise_tensor
batch_crosswise_diffs = crosswise_tensor(
train_features,
train_features,
batch_indices,
batch_nn_indices,
)
from MuyGPyS.gp.tensors import pairwise_tensor
pairwise_diffs = pairwise_tensor(
train_features, batch_nn_indices
)
Kcross = muygps.kernel(batch_crosswise_diffs)
K = muygps.kernel(pairwise_diffs)
batch_targets = train_responses[batch_indices, :]
batch_nn_targets = train_responses[batch_nn_indices, :]
from MuyGPyS.gp.tensors import make_train_tensors
(
batch_crosswise_diffs,
batch_pairwise_diffs,
batch_targets,
batch_nn_targets,
) = make_train_tensors(
batch_indices,
batch_nn_indices,
train_features,
train_responses,
)
from MuyGPyS.optimize import optimize_from_tensors
muygps = optimize_from_tensors(
muygps,
batch_targets,
batch_nn_targets,
batch_crosswise_diffs,
batch_pairwise_diffs,
loss_method="lool",
obj_method="loo_crossval",
opt_method="bayesian",
verbose=False,
random_state=1,
init_points=5,
n_iter=2,
)
from MuyGPyS.optimize.sigma_sq import muygps_sigma_sq_optim
K = muygps.kernel(batch_pairwise_diffs)
muygps = muygps_sigma_sq_optim(muygps, batch_pairwise_diffs, batch_nn_targets, sigma_method="analytic")
Matplotlib is building the font cache; this may take a moment.
Fast Prediction
With set (or learned) hyperparameters, we are able to use the muygps object for fast prediction capability. Several workflows are supported.
See below a fast posterior mean workflow, using the data structures built up in this example. This workflow uses the compact tensor-making function make_fast_predict_tensors() to succinctly create tensors defining the pairwise_diffs among each nearest neighbor and the train_nn_targets_fast or responses of the nearest neighbors in each set. We then create theK covariance tensor and form the precomputed coefficients matrix. We then pass the
precomputed coefficients matrix, the updated nn_indices matrix, and the closest neighbor of each test point to MuyGPS.fast_posterior_mean() in order to obtain our predictions.
[3]:
from MuyGPyS.gp.tensors import make_fast_predict_tensors, fast_nn_update, make_predict_tensors
nn_indices,_ = nbrs_lookup.get_nns(train_features)
nn_indices = nn_indices.astype(int)
[4]:
from MuyGPyS.gp.tensors import make_predict_tensors
nn_indices = fast_nn_update(nn_indices)
#find the closest training point to each test point, and its corresponding nearest neighbor set
test_neighbors, _ = nbrs_lookup.get_nns(test_features)
closest_neighbor = test_neighbors[:, 0]
closest_set = nn_indices[closest_neighbor, :].astype(int)
#make crosswise distances tensor for prediction
(
crosswise_diffs,
pairwise_diffs,
nn_targets,
) = make_predict_tensors(
np.arange(test_count),
closest_set,
test_features,
train_features,
train_responses,
)
K = muygps.kernel(pairwise_diffs)
precomputed_coefficients_matrix = muygps.fast_coefficients(
K,
nn_targets)
#perform GP regression
Kcross = muygps.kernel(crosswise_diffs)
fast_predictions = muygps.fast_posterior_mean(
Kcross,
precomputed_coefficients_matrix[closest_neighbor,:,:])
Regular Prediction
With set (or learned) hyperparameters, we are able to use the muygps object to predict the response of test data. Several workflows are supported.
See below a simple posterior mean workflow, using the data structures built up in this example. This workflow uses the compact tensor-making function make_predict_tensors() to succinctly create tensors defining the pairwise_diffs among each nearest neighbor set, the crosswise_diffs between each test point and its nearest neighbor set, and the nn_targets or responses of the nearest neighbors in each set. We then create the Kcross cross-covariance
matrix and K covariance tensor and pass them to MuyGPS.posterior_mean() in order to obtain our predictions.
[5]:
from MuyGPyS.gp.tensors import make_predict_tensors
# make the indices
test_count, _ = test_features.shape
indices = np.arange(test_count)
nn_indices, _ = nbrs_lookup.get_nns(test_features)
# make difference and target tensors
(
crosswise_diffs,
pairwise_diffs,
nn_targets,
) = make_predict_tensors(
indices,
nn_indices,
test_features,
train_features,
train_responses,
)
# Make the kernel
Kcross = muygps.kernel(crosswise_diffs)
K = muygps.kernel(pairwise_diffs)
# perform Gaussian process regression
predictions = muygps.posterior_mean(
K, Kcross, nn_targets
)
Timing Experiment
We compare the prediction time of a regular posterior mean workflow to that of the fast posterior mean workflow. In the regular posterior mean workflow we compute the sum of the time it takes to identify the nearest neighbors of the test features, the time it takes to form the relevant kernel tensors, and the time to solve the posterior means. In the fast posterior mean case, we compute the sum of the time it takes to identify the nearest neighbor of each test point, the coefficient lookup in the precomputed coefficient matrix, and the dot product to form posterior means.
[6]:
from MuyGPyS.optimize.loss import mse_fn
import timeit
test_count, _ = test_features.shape
indices = np.arange(test_count)
def timing_posterior_mean():
nn_indices, _ = nbrs_lookup.get_nns(test_features)
(
crosswise_diffs,
pairwise_diffs,
nn_targets,
) = make_predict_tensors(
indices,
nn_indices,
test_features,
train_features,
train_responses,
)
Kcross = muygps.kernel(crosswise_diffs)
K = muygps.kernel(pairwise_diffs)
predictions = muygps.posterior_mean(
K, Kcross, nn_targets
)
print(f"regular RMSE:")
print(f"\tRMSE: {np.sqrt(mse_fn(predictions, test_responses))}")
print("regular prediction time:")
%timeit timing_posterior_mean()
nn_indices = fast_nn_update(nn_indices)
def timing_fast_posterior_mean():
test_neighbors, _ = nbrs_lookup.get_nns(test_features)
closest_neighbor = test_neighbors[:, 0]
closest_set = nn_indices[closest_neighbor, :].astype(int)
#make crosswise distances tensor
(
crosswise_diffs,
pairwise_diffs,
nn_targets,
) = make_predict_tensors(
np.arange(test_count),
closest_set,
test_features,
train_features,
train_responses,
)
Kcross = muygps.kernel(crosswise_diffs)
fast_predictions = muygps.fast_posterior_mean(
Kcross,
precomputed_coefficients_matrix[closest_neighbor,:,:])
print(f"fast prediction RMSE:")
print(f"\tRMSE: {np.sqrt(mse_fn(fast_predictions, test_responses))}")
print("fast prediction time:")
%timeit timing_fast_posterior_mean()
regular RMSE:
RMSE: 0.1206710068099257
regular prediction time:
64.7 ms ± 1.92 ms per loop (mean ± std. dev. of 7 runs, 10 loops each)
fast prediction RMSE:
RMSE: 1.658329619936535
fast prediction time:
14.8 ms ± 101 µs per loop (mean ± std. dev. of 7 runs, 100 loops each)
Results
We achieve roughly two orders of magnitude speedup using the fast prediction acceleration. The improvement is even more dramatic when the methods are implemented in JAX.